Gene2Phenotype: A Database of Structured Human Monogenic Diseases and Pathomechanisms
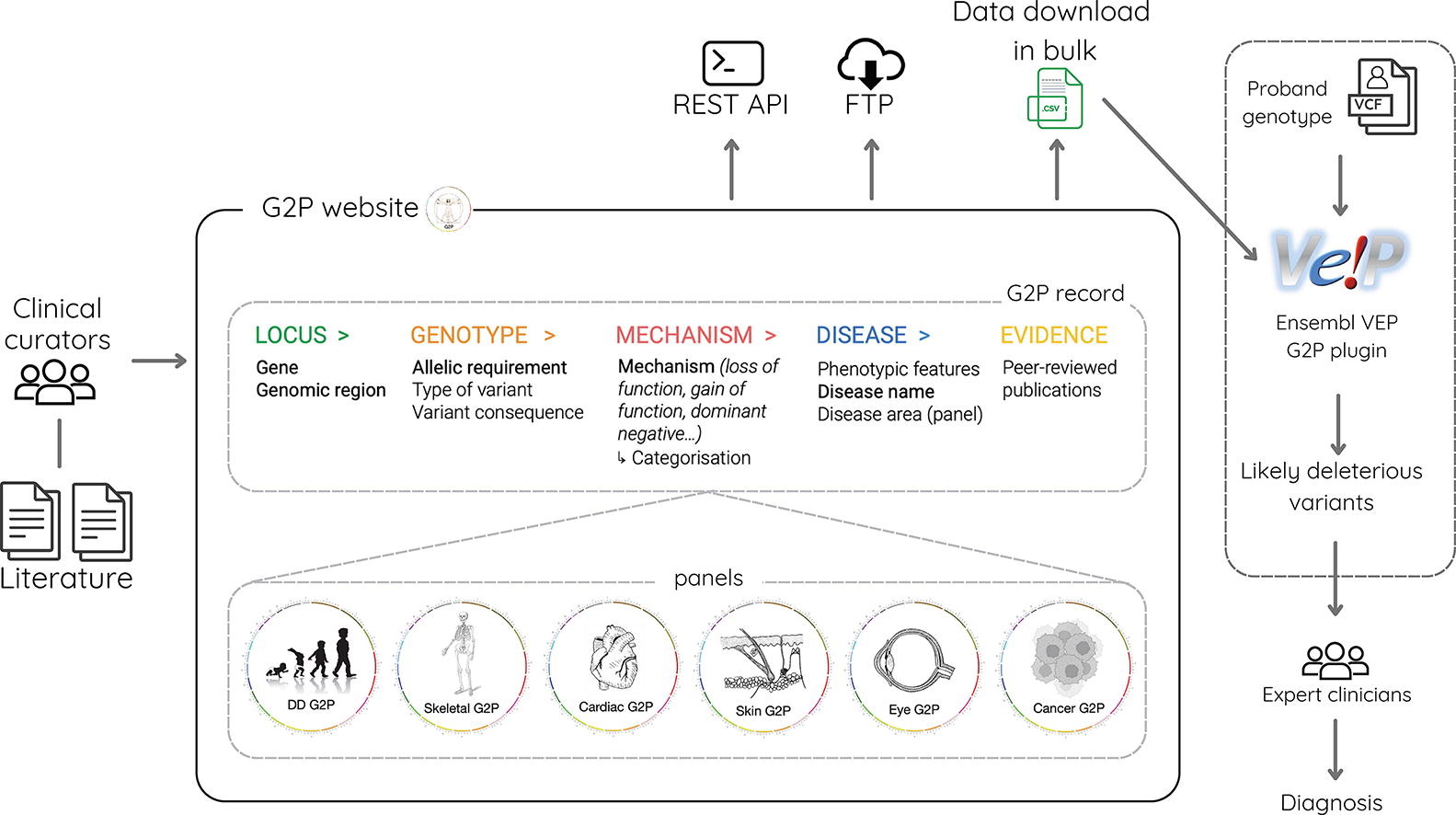
Gene2Phenotype (G2P) is an open, expert-curated database designed to make monogenic (single-gene) disease knowledge more accessible, computable, and clinically useful. Originally created in 2012 to support variant filtering in the Deciphering Developmental Disorders study, G2P has expanded beyond developmental disorders to cover additional domains including cardiac, eye, ear, skeletal, skin disorders, and germline cancer predisposition. This paper describes a major redesign of both the data model and the platform, motivated by the growing recognition that gene–disease links are not sufficiently described by “gene X causes disease Y”—instead, downstream applications increasingly need structured details about inheritance, genotype patterns, and especially the molecular pathomechanism (e.g., loss of function vs gain of function vs dominant negative).
At the core of the updated resource is a structured “G2P record” built around a modular Locus–Genotype–Mechanism–Disease–Evidence (LGMDE) thread. Each record captures allelic requirement (e.g., monoallelic/biallelic, de novo, incomplete penetrance), high-level and more specific mechanism categories, observed variant types and inferred variant consequences (using standard ontologies), linked phenotypes (HPO terms), and curated supporting evidence from the literature. The authors report that G2P currently contains 3828 records across 7 partially overlapping panels, with most records rated moderate-to-definitive confidence and many cross-referenced to OMIM and Mondo. Alongside the model refresh, the team delivered a redesigned website for exploration plus a public REST API and bulk downloads (including monthly frozen snapshots), aiming to improve FAIRness and simplify integration into clinical and research workflows.
A key applied contribution is continued support for diagnostic variant filtering via the VEP-G2P plugin for Ensembl’s Variant Effect Predictor, which matches patient genotypes to curated allelic requirements and (optionally) to expected molecular consequences, using “strict” or “tolerant” consequence matching modes. The paper also situates G2P within a broader ecosystem (e.g., GenCC, PanelApp, ClinGen, OMIM, Open Targets, Ensembl/UCSC), arguing that G2P’s distinctive value is its consistently structured representation of disease mechanism and variant consequence—information that can improve variant prioritisation, guide assay relevance (e.g., MAVEs), and support precision medicine and therapeutic development. The authors note limitations in coverage and incomplete backfilling of newer annotations into older records, and outline future plans to use NLP and machine learning for literature surveillance and curation “seeding” while clearly distinguishing mined vs curator-validated content.
For more information the full article is published here:
Sarah E. Hunt, Diana Lemos, Seeta Ramaraju Pericherla, Olanrewaju Austine-Orimoloye, Elena Cibrian Uhalte, T. Michael Yates, Morad Ansari, Louise Thompson, Julia Foreman, T. Ian Simpson, James S. Ware, Caroline F. Wright, Mallory A. Freeberg, Helen V. Firth. Gene2Phenotype: A Database of Structured Human Monogenic Diseases and Pathomechanisms. Journal of Molecular Biology, 2026,169768. https://doi.org/10.1016/j.jmb.2026.169768